Olga A. Omelchuk,a,b@1 Nikita M. Belov,а Vladimir B. Tsvetkov,c,d,e Natalia E. Grammatikova,a,f Lyudmila N. Lysenkova,a Alexander M. Korolev,a Olga B. Bekker,g Valery N. Danilenko,g and Andrey E. Shchekotikhina,b@2
aGause Institute of New Antibiotics, 119021 Moscow, Russian Federation
bD. I. Mendeleev University of Chemical Technology of Russia, 125047 Moscow, Russian Federation
cTopchiev Institute of Petrochemical Synthesis, Russian Academy of Sciences, 119991 Moscow, Russian Federation
dInstitute for Physical-Chemical Medicine, 119435 Moscow, Russian Federation
eResearch Institute of Influenza, 197376 St.-Petersburg, Russian Federation
fI.M. Sechenov First Moscow State Medical University, 119991 Moscow, Russian Federation
gN.I. Vavilov Institute of General Genetics, Russian Academy of Sciences, 119333 Moscow, Russian Federation
@1Corresponding author E-mail: omelchuk.93@mail.ru
@2Corresponding author E-mail: shchekotikhin@mail.ru
DOI: 10.6060/mhc160964o
Macroheterocycles 2016 9(4) 453-461
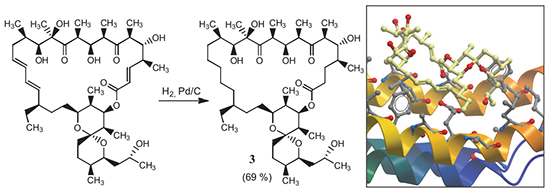
Oligomycin A (1) belongs to the class of highly substituted macrolide antibiotics produced by actinomycetes Streptomyces.[1] Oligomycin A (1) has antifungal and cytotoxic activities but Gram-negative and Gram-positive bacteria are resistant to 1 except actinobacteria.[2] At micromolar concentrations 1 inhibited F0F1-ATPase by binding with subunit F0 and blocking OSCP (oligomycin sensitivity conferring protein).[3] Previously a series of modifications at the side chain and chemical transformation of the lactone moiety of 1 have been developed.[4-8] However, all derivatives modificated at the lactone moiety were less potent than 1.[7,8]Recently a new derivative, 2,3-dihydrooligomycin A (2), has been obtained by a microbiological method. This compound showed a higher activity against Saccharomyces cerevisiae than 1.[9] We investigated chemical reduction of double C-C bonds of 1. Hydrogenation of 1 on Pd/BaSO4 gives an unseparatable mixture of products. Using Pd/C in the reduction of 1 in methanol led to 2,3,16,17,18,19-hexahydrooligomycin A (3) in a good yield. The structure of 3 was confirmed by NMR-spectroscopy and high resolution mass spectrometry. Investigations of antifungal and anti-yeast properties were evaluated against Candida spp. (clinical isolates and reference strains) and filamentous fungi. Hexahydrooligomycin (3) showed a high selective activity against C. parapsilosis, C. utilis, C. tropicalis, and C. krusei and was inefficient against micromycetes. Notably, the C. krusei strain is resistant to fluconazole. The actinomyces S. fradiae ATCC-19069 exceptionally sensitive to 1 (MIC 0.5 nmol/ml) was resistant to 3. Hexahydrooligomycin 3 was also less toxic than 1 against human K562 leukemia and HCT116 colon carcinoma cell lines. Thus, reduction of three double C=C bonds increases selectivity to Candida spp. with simultaneous decrease of potency against other test-cultures. A computer assisted simulation of interaction of 1 and 3 with the intracellular target (F0-subunit of the ATP-synthase) was performed using Molsoft ICM-Pro 3.8. The results of docking studies were in a good agreement with bioscreening data. Further development of methods of modification of 1 is needed for detailed understanding of SAR and drug optimization.
References: